Introduction
Support vector machine (SVM) is a non-linear classifier which is often reported as producing superior classification results compared to other methods. The idea behind the method is to non-linearly map the input data to some high dimensional space, where the data can be linearly separated, thus providing great classification (or regression) performance. One of the bottlenecks of the SVM is the large number of support vectors used from the training set to perform classification (regression) tasks. In my code, I use SSE optimization to increase performance.
Background (optional)
www.kernel-machines.org is a great source for SVM information.
Using the Code
In SVM class, I use my 2D SSE optimized vector code for faster computation. The SVMachine class contains the following functions you need to use:
SVMachine::SVMachine(const wchar_t* fname); ctor int SVMachine::status() const; status after ctor (0 upon success and negative in case of errors) unsigned int SVMachine::dimension() const; the dimensionality of the SVM int SVMachine::classify(const float* x, double& y) const; to classify unknown vector x
ctor reads SVM configuration from file having the following text format:
input vector dimensionality
number of support vectors
kernel type [kernel parameter]
bias
weight
1st support vector
weight
2nd support vector
...
For example, polynomial kernel SVM for iris data set to classify setosa from virgi consisted from 4 support vectors is presented below:
4
4
polynomial 3
1.1854890124462447
7.792329562624775e-012
51
33
17
5
9.9563612529691003e-012
48
34
19
2
1.0631195782759572e-011
51
38
19
4
-2.8372847134273557e-011
49
25
45
17
The SVM decision function is presented by the following formula:
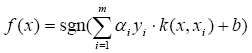
Where x is the input vector, alpha and y are the weights of the support vectors, having y as positive or negative class mark (+1 or -1) and b is the bias. From the iris SVM file, we can see that there are 4 four dimensional support vectors (3 first from positive class being setosa samples and the last one from negative class pertaining to virgi), the kernel is the polynomial one with 3 as the parameter, the bias is equal to 1.1854890124462447.
In my class, I use 3 kernels.
linear:
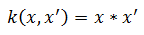
rbf:
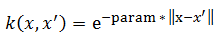
polynomial:
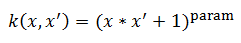
Where param is the [kernel parameter] in the SVM file.
Typically a grid search is used to select best classification (regression) results by varying alpha and param over some range. In my iris SVM, the alphas are equal to 1 and param is the degree of polynomial.
To classify unknown vector sample x as belonging to positive or negative class, use SVMachine::classify() function. It returns +1 or -1 as the result of classification and provides to y the result of sum from the SVM decision formula.
Points of Interest
Add other kernels.
History
- 14th April, 2008: Initial post
Highly skilled Engineer with 14 years of experience in academia, R&D and commercial product development supporting full software life-cycle from idea to implementation and further support. During my academic career I was able to succeed in MIT Computers in Cardiology 2006 international challenge, as a R&D and SW engineer gain CodeProject MVP, find algorithmic solutions to quickly resolve tough customer problems to pass product requirements in tight deadlines. My key areas of expertise involve Object-Oriented
Analysis and Design OOAD, OOP, machine learning, natural language processing, face recognition, computer vision and image processing, wavelet analysis, digital signal processing in cardiology.
 General
General  News
News  Suggestion
Suggestion  Question
Question  Bug
Bug  Answer
Answer  Joke
Joke  Praise
Praise  Rant
Rant  Admin
Admin